Understanding the origin and genetic basis of speciation and adaptation is the overarching theme and motivation of my research program in evolutionary biology. In an integrative approach, my group and I combine population genomics empowered by next-generation sequencing data with classical morphometric and ecological study and experiments. Our aim is to unravel how the origin of genetic variation and genomic architecture may constrain or facilitate adaptation and speciation and thereby govern the speed and predictability of both processes. By answering fundamental questions about the origin of biodiversity, we hope to contribute to better understanding and alleviating the current biodiversity crisis as well as predicting its consequences for the future.
On the genomic origin of large gull species
Large white-headed gulls (Larus spp.) are one of the fastest diversifying bird lineages with 16 species and 36 subspecies distributed mostly around the Northern hemisphere going back to a common ancestor approx. 1 million years ago. Many of these taxa are still in the speciation process and as lineage, gulls span the entire speciation continuum from diverging populations to sympatric, fully isolated species. In our new research project funded by the Swiss National Science Foundation, we are going to generate state-of-the-art genomic resources to understand what genomic changes are associated with the speciation continuum in this fantastic and confusing bird lineage. Our aim is to investigate the role of genome structure evolution and combinatorial mechanisms (Marques et al. 2019) facilitating rapid speciation and to unravel how the genomic architecture of reproductive isolation changes across the speciation continuum. This project is a collaboration with Walter Salzburger, Joana Meier and the Wellcome Sanger Institute, Pierre-André Crochet, Magdalena Zagalska-Neubauer and Manuel Schweizer.

Unweaving the largest vertebrate adaptive radiation
The East African Lake Victoria contains more than 500 species of cichlid fish that evolved since the lake re-filled 15,000 years ago, making this the largest young adaptive radiation among vertebrates. My former research as Postdoc in the lab of Ole Seehausen at University of Bern and Eawag, Switzerland, in collaboration with Joana Meier, Matt McGee and Laurent Excoffier, used the power of whole genome re-sequencing data to reconstruct the evolutionary history behind this evolutionary phenomenon. We used >500 re-sequenced genomes of >250 species from Lake Victoria and replicate radiations in the Lake Victoria region, which were seeded ~150,000 years ago by a hybrid swarm formed by two 1.5 million years divergent cichlid lineages from the Nile and the Congo drainage (Meier et al. 2017). Our aim was to unravel the genomic mechanisms underlying repeated evolution of ecomorphs in replicate radiations, to test different hypotheses on the origin of adaptive radiations (McGee et al. 2020), such as constraints of genetic variation and the role of a hybrid swarm in facilitating radiations (Marques et al. 2019), and to reconstruct the timing and order of adaptive radiations. I co-supervised three fantastic MSc students, Vera Ogi, Mikki Law and Mark Charran in this project who worked on genomic and morphometric aspects.

Adaptation genomics of a stickleback radiation
The adaptive radiation of threespine stickleback on the Haida Gwaii archipelago in Western Canada started after the retreat of the ice sheets roughly 12,000 years ago, where stickleback invaded many different ecological theaters, varying in predator composition, water chemistry and system size from large to small lakes and streams, oligotrophic to dystrophic to eutorphic lakes. Adaptation in stickelback to these environments led to the largest phenotypic diversity observed among threespine stickleback. Tom Reimchen identified light spectrum, system size and predator composition as the main predictors of phenotypic variation. In my postdoc with Tom at the University of Victoria, Canada, and in collaboration with David Kingsley, Felicity Jones and Federica di Palma, I have been exploring the genomic targets of repeated selection across the radiation using 58 whole genomes and a selection experiment. We discovered that adaptation can be highly predictable at the molecular level, both from standing genetic variation on short time scales such as a 19-year selection experiment (Marques et al. 2018) or in the 12,000 year old adaptive radiation, and across 200 million years, where we found convergent molecular adaptation of a color vision gene to the same blackwater habitat in stickleback and distantly related fish species (Marques et al. 2017). Signatures of parallel genomic adaptation across the radiation further suggested that the more distant a new ecological niche is from the original habitat, the more genetic loci are under selection (Marques et al. 2022).


Genomics of rapid ecological speciation
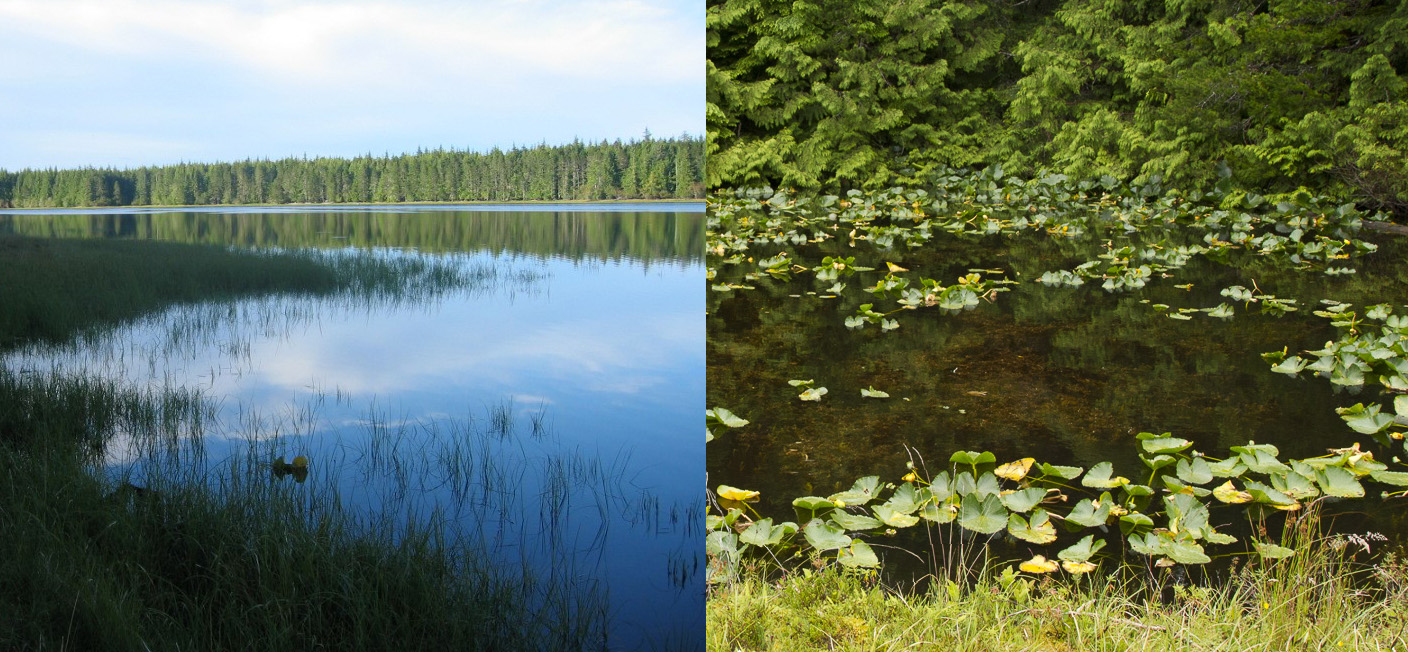
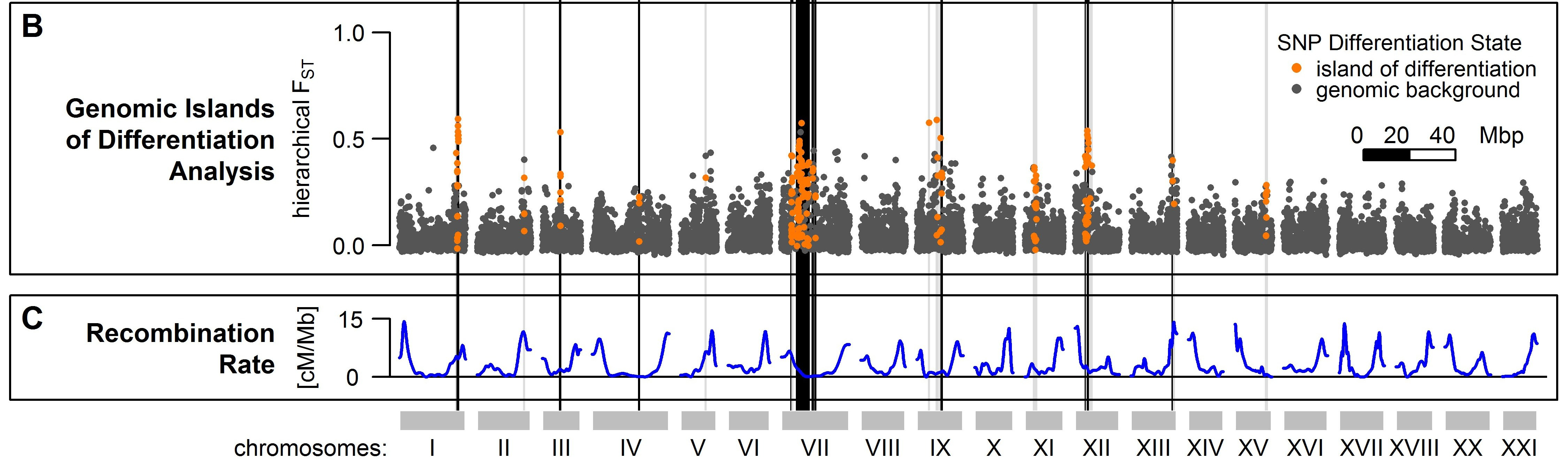
Threespine stickleback have been released into the large Lake Constance catchment around 150 years ago and have since then colonized both lake and stream habitats successfully, making roughly 50% of today’s fish biomass in the lake. Adaptation to the two habitats led to divergence between lake and stream ecotypes in body size, body armor, feeding morphology, nuptial color and life history: lake fish get much bigger, have larger bony plates, eat more planctonic than benthic prey, have redder throats, start breeding one year later and get older than stream fish. During my PhD research supervised by Ole Seehausen, Laurent Excoffier and Katie Wagner and conducted at the University of Bern and Eawag, Switzerland, in close collaboration with my colleagues Kay Lucek and Joana Meier, we showed ecotypes in Lake Constance breed in sympatry while maintaining ecotypic divergence, making them an unusual case of ecological speciation with very early sympatry. Genomic divergence associated with these two ecotypes persists on multiple chromosomes despite ongoing gene flow, in particular in low recombination regions where the genetic basis of many divergent traits is also found (Marques et al. 2016).

Speciation driven by selection on nuptial color
A very young (<90 years) old pond near Bern harbors a small, peculiar population of threespine stickleback with two male morphs: larger red-throated males breed on the steep and shady shore and make tiny, concealed nests, while smaller orange-throated males breed on the open, sunny and deeper pond bottom and make large crater nests. My second PhD project in the Seehausen and Excoffier labs, in close collaboration with my colleague Kay Lucek and BSc student Anna Feller who I co-supervised with Ole, showed that divergence is limited to nuptial coloration and extended phenotype (nest), but absent in predator defense, feeding and swimming performance traits, and that this divergence is reflected in moderate genomic divergence on multiple chromosomes containing potential candidate genes involved in color vision (Marques et al. 2016). We also found that femaes prefer either the orange males or show a tendency to prefering the red male in an experimental setup (Feller et al. 2017). Therefore, sexual selection, interacting with the environment, has likely led to divergence and stabilization of these two morphs in full sympatry – a situation that could be the earliest stage of ecological speciation.
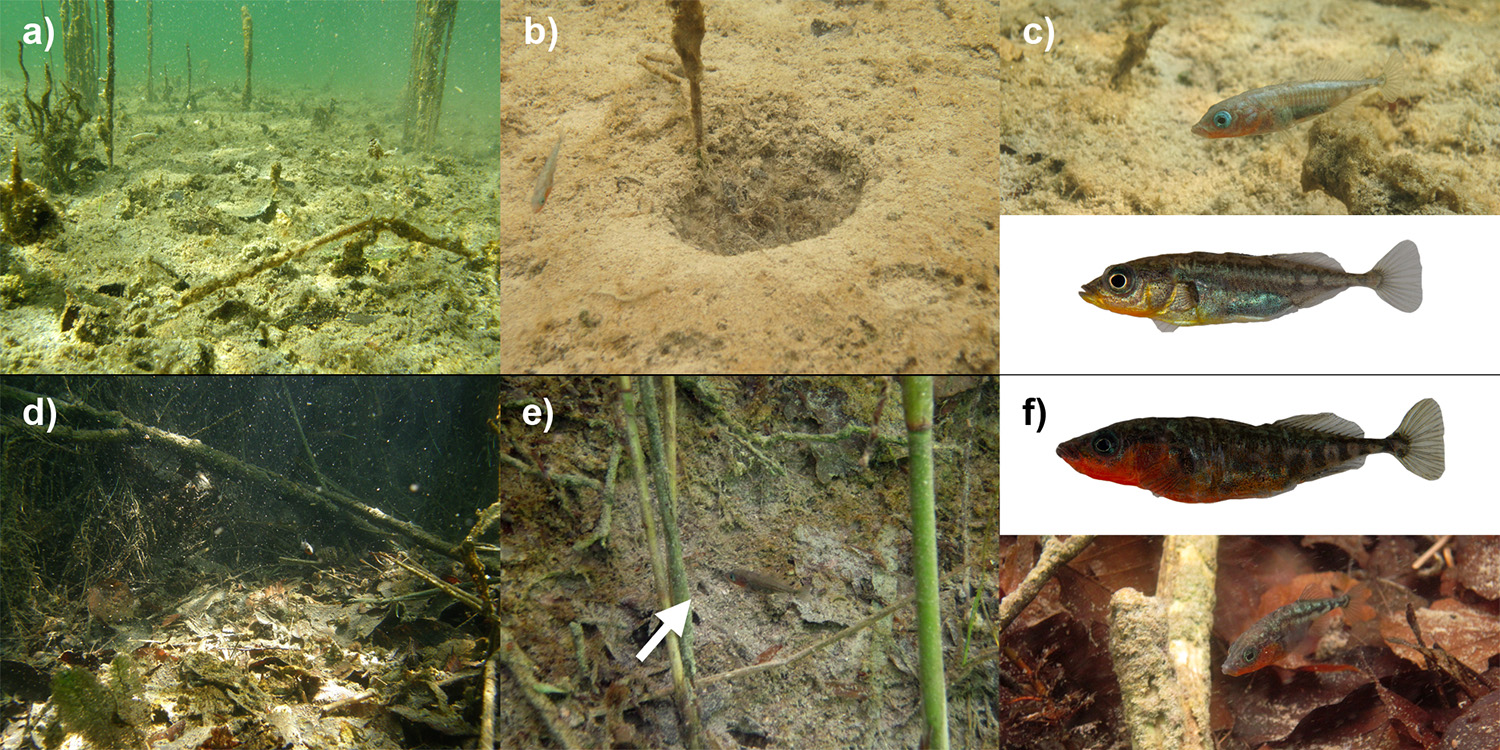

Origin of alleles in an emerging adaptive radiation
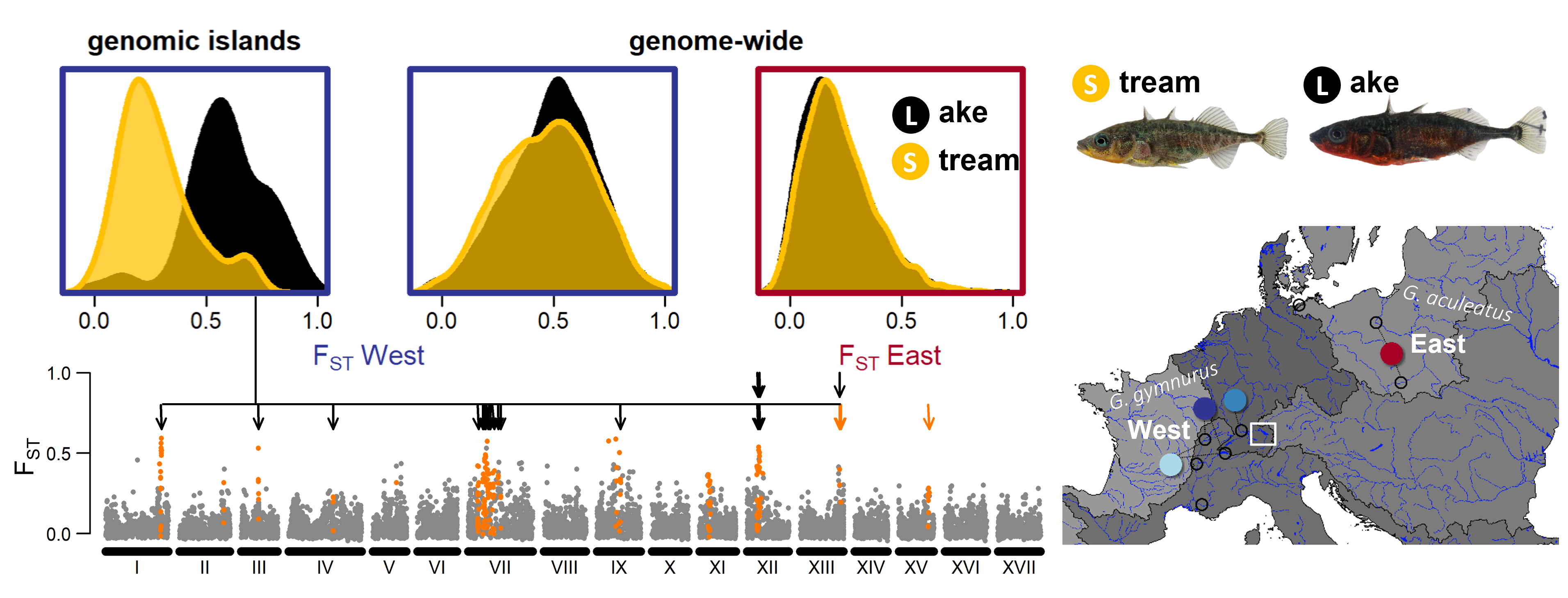
Stickleback from very different origin have adapted to lake and stream environments in several catchments across Switzerland: a Rhone lineage from South-Western Europe was introduced into Lake Geneva, an Eastern European lineage into Lake Constance and a mixture of these two and a Northern European Rhine lineage have colonized Lake Biel. My PhD research showed that ecotype differentiation in the genome persist only in Lake Constance against high amounts of gene flow. Demographic modelling allowed us to date the split between the different lineages colonizing these lakes and to find out that gene flow between the lineages has occurred and likely contributed to the repeated divergence of lake and stream ecotypes (Marques et al. 2019, Marques et al. 2020).

You must be logged in to post a comment.